Code
library(data.table)
library(MSstats)
library(qs2)
library(dplyr)
library(ggplot2)
library(ggrepel)
library(tidyverse)
library(readr)base_dir <- "/nemo/lab/destrooperb/home/shared/zanettc/giulia_proteomics/bulk_proteomics"
run_num <- "run4"
results_dir <- file.path(base_dir, "results", run_num)
objects_dir <- file.path(base_dir, "data", "processed", run_num)
input_dir <- file.path(base_dir, "input")
dir.create(results_dir, recursive = TRUE, showWarnings = FALSE)
dir.create(objects_dir, recursive = TRUE, showWarnings = FALSE)All 16 samples passed QC. R.Condition = “Not Defined” in the raw file because the study design was not configured inside Spectronaut — we define it here from the known design.
Only the columns required by MSstats are read — this keeps memory manageable on the 8.4 GB file.
msstats_cols <- c(
"R.Condition", "R.FileName", "R.Replicate",
"PG.ProteinGroups", "PG.ProteinAccessions",
"PEP.StrippedSequence",
"EG.ModifiedSequence", "EG.PrecursorId", "EG.Qvalue",
"FG.Charge", "FG.Id",
"F.FrgIon", "F.Charge", "F.FrgLossType", "F.FrgNum", "F.FrgType",
"F.NormalizedPeakArea", "F.PeakArea", "F.ExcludedFromQuantification"
)
cat("Reading Spectronaut MSstats report...\n")Reading Spectronaut MSstats report...Rows loaded: 24057952 SpectronauttoMSstatsFormat uses R.Condition internally before merging with the annotation. We replace “Not Defined” with the correct values from the annotation to avoid any issues.
Condition distribution after fix: R.Condition R.FileName N
<char> <char> <int>
1: hApp GA_1 1503622
2: hApp GA_2 1503622
3: hApp GA_3 1503622
4: hApp GA_4 1503622
5: hAppNLGF GA_5 1503622
6: hAppNLGF GA_6 1503622
7: hAppNLGF GA_7 1503622
8: hAppNLGF GA_8 1503622
9: hAppNLGF_FIRE GA_10 1503622
10: hAppNLGF_FIRE GA_11 1503622
11: hAppNLGF_FIRE GA_12 1503622
12: hAppNLGF_FIRE GA_9 1503622
13: hApp_FIRE GA_13 1503622
14: hApp_FIRE GA_14 1503622
15: hApp_FIRE GA_15 1503622
16: hApp_FIRE GA_16 1503622Confirms that hApp and hAppNLGF conditions have detectable Aβ signal. In microglia, signal reflects phagocytosed / secreted Aβ rather than endogenous APP expression.
arctic_peptide <- "LVFFAGDVGSNK"
normal_peptide <- "LVFFAEDVGSNK"
peptide_both <- "DAEFRHDSGYEVHHQK"
target_peptide <- "LVFFAEDVGSNK" # Conserved Aβ tryptic peptide (mouse and human APP)
ab_check <- raw_spectronaut %>%
filter(PEP.StrippedSequence == target_peptide) %>%
group_by(R.FileName, R.Condition) %>%
summarise(Total_Intensity = sum(F.PeakArea, na.rm = TRUE), .groups = "drop") %>%
mutate(Log2_Intensity = log2(Total_Intensity + 1)) %>%
# Extract numeric sample ID for proper sorting
mutate(Sample_Num = as.numeric(gsub("GA_", "", R.FileName))) %>%
arrange(R.Condition, Sample_Num) %>%
mutate(R.FileName = factor(R.FileName, levels = unique(R.FileName)))
if (nrow(ab_check) == 0) {
cat("Aβ peptide (", target_peptide, ") not detected in any sample.\n")
} else {
cat("Aβ peptide detected in", nrow(ab_check), "sample(s):\n")
print(ab_check)
}Aβ peptide detected in 16 sample(s):
# A tibble: 16 × 5
R.FileName R.Condition Total_Intensity Log2_Intensity Sample_Num
<fct> <chr> <dbl> <dbl> <dbl>
1 GA_1 hApp 3870. 11.9 1
2 GA_2 hApp 5470. 12.4 2
3 GA_3 hApp 2421. 11.2 3
4 GA_4 hApp 2905. 11.5 4
5 GA_5 hAppNLGF 25.4 4.72 5
6 GA_6 hAppNLGF 93.3 6.56 6
7 GA_7 hAppNLGF 25.3 4.72 7
8 GA_8 hAppNLGF 12.4 3.75 8
9 GA_9 hAppNLGF_FIRE 72.5 6.20 9
10 GA_10 hAppNLGF_FIRE 16.4 4.12 10
11 GA_11 hAppNLGF_FIRE 8.20 3.20 11
12 GA_12 hAppNLGF_FIRE 99.1 6.64 12
13 GA_13 hApp_FIRE 3174. 11.6 13
14 GA_14 hApp_FIRE 2566. 11.3 14
15 GA_15 hApp_FIRE 2386. 11.2 15
16 GA_16 hApp_FIRE 3082. 11.6 16p_ab <- ggplot(ab_check, aes(x = R.FileName, y = Log2_Intensity, fill = R.Condition)) +
geom_bar(stat = "identity") +
scale_fill_manual(values = c(
"hApp" = "#1b9e77", "hAppNLGF" = "#d95f02",
"hAppNLGF_FIRE" = "#7570b3", "hApp_FIRE" = "#e7298a"
)) +
theme_minimal() +
theme(axis.text.x = element_text(angle = 45, hjust = 1)) +
labs(title = "Aβ Tryptic Peptide Signal per Sample",
subtitle = paste0("Peptide: ", target_peptide),
x = "Sample", y = "Log2 Intensity")
ggsave(file.path(results_dir, "AB_Peptide_Verification.pdf"), p_ab, width = 10, height = 6)
print(p_ab)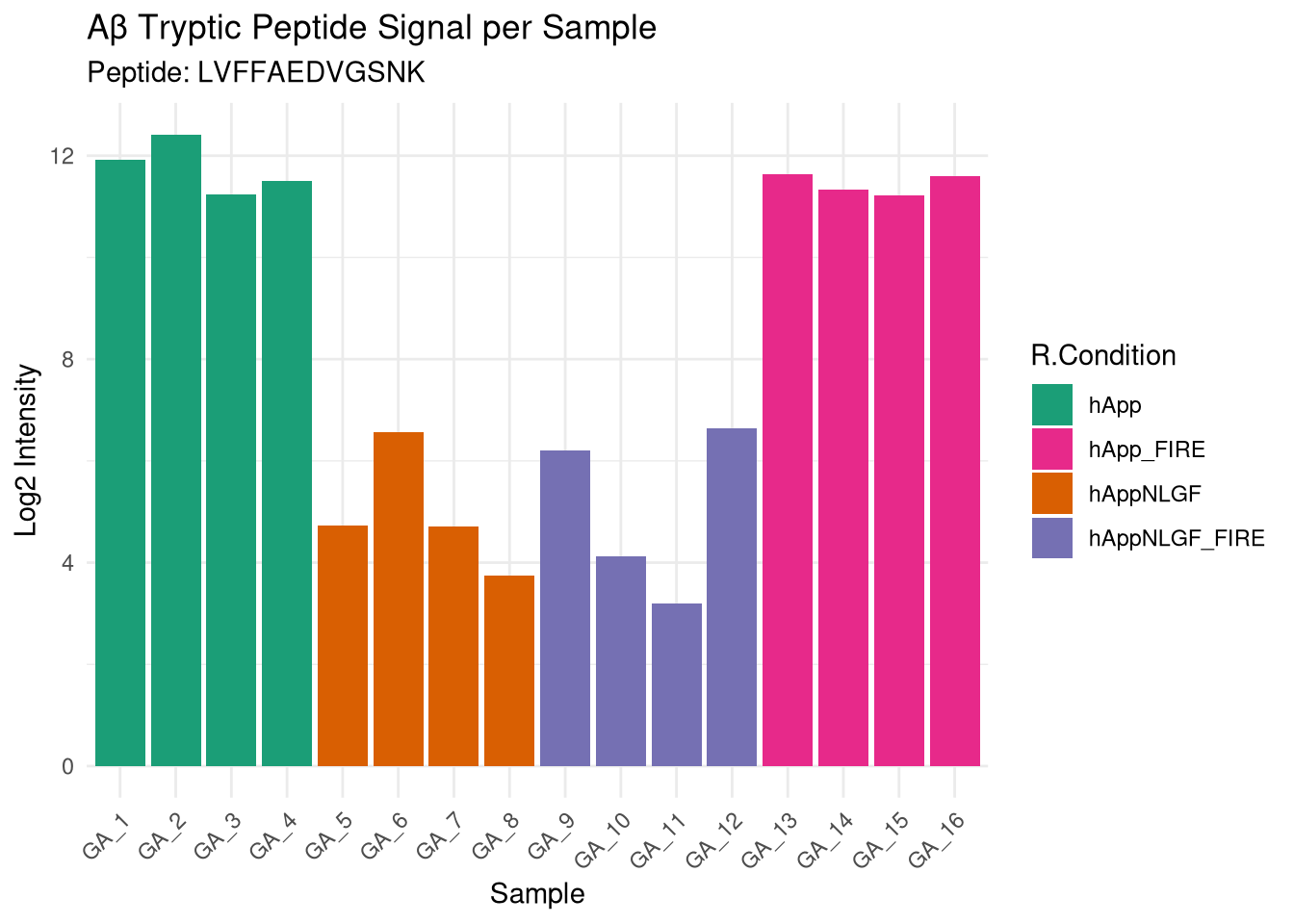
Feature-level filtering: Qvalue < 0.01, remove features with too few measurements.
INFO [2026-04-29 22:21:08] ** Raw data from Spectronaut imported successfully.
INFO [2026-04-29 22:21:14] ** Raw data from Spectronaut cleaned successfully.
INFO [2026-04-29 22:21:14] ** Using provided annotation.
INFO [2026-04-29 22:21:14] ** Run labels were standardized to remove symbols such as '.' or '%'.
INFO [2026-04-29 22:21:14] ** The following options are used:
- Features will be defined by the columns: PeptideSequence, PrecursorCharge, FragmentIon, ProductCharge
- Shared peptides will be removed.
- Proteins with single feature will not be removed.
- Features with less than 3 measurements across runs will be removed.
INFO [2026-04-29 22:21:20] ** Intensities with values of FExcludedFromQuantification equal to TRUE are replaced with NA
WARN [2026-04-29 22:21:23] ** PGQvalue not found in input columns.
INFO [2026-04-29 22:21:24] ** Intensities with values not smaller than 0.01 in EGQvalue are replaced with NA
INFO [2026-04-29 22:21:30] ** Features with all missing measurements across runs are removed.
INFO [2026-04-29 22:21:36] ** Shared peptides are removed.
INFO [2026-04-29 22:22:56] ** Multiple measurements in a feature and a run are summarized by summaryforMultipleRows: max
INFO [2026-04-29 22:23:00] ** Features with one or two measurements across runs are removed.
INFO [2026-04-29 22:23:02] ** Run annotation merged with quantification data.
INFO [2026-04-29 22:23:16] ** Features with one or two measurements across runs are removed.
INFO [2026-04-29 22:23:16] ** Fractionation handled.
INFO [2026-04-29 22:23:52] ** Updated quantification data to make balanced design. Missing values are marked by NA
INFO [2026-04-29 22:23:55] ** Finished preprocessing. The dataset is ready to be processed by the dataProcess function.MSstats input rows: 11946400 Unique proteins: 8918 Keeps features observed in ≥50% of runs within at least one condition. This removes near-absent features that destabilise the linear model. This is the same as saying proteins must be present in at least 2 samples out of 3 for at least one condition.
msstats_input_filt <- as.data.frame(msstats_input) %>%
group_by(ProteinName, PeptideSequence, PrecursorCharge, FragmentIon, ProductCharge, Condition) %>%
summarise(
n_obs = sum(!is.na(Intensity) & Intensity > 0),
n_total = n(),
prop = n_obs / n_total,
.groups = "drop"
) %>%
group_by(ProteinName, PeptideSequence, PrecursorCharge, FragmentIon, ProductCharge) %>%
filter(any(prop >= 0.5)) %>%
ungroup() %>%
inner_join(
as.data.frame(msstats_input),
by = c("ProteinName", "PeptideSequence", "PrecursorCharge",
"FragmentIon", "ProductCharge", "Condition")
)
message("Rows before completeness filter: ", nrow(msstats_input))
message("Rows after completeness filter: ", nrow(msstats_input_filt))
message("Proteins retained: ", uniqueN(msstats_input_filt$ProteinName))
table(
Value_Type = case_when(
is.na(msstats_input$Intensity) ~ "True NAs",
msstats_input$Intensity == 0 ~ "Exact Zeros (Filtered by Q-value)",
msstats_input$Intensity > 0 ~ "Valid Measurements"
)
)Value_Type
True NAs Valid Measurements
1736082 10210318 The MSstats report contains only UniProt accessions in ProteinName — no gene symbols. To filter keratins, build a lookup from UniProt or biomaRt and enable the block below.
## Keratin protein filter
protein_dict <- read_csv(file.path(base_dir, "data", "processed", "run1", "protein_dictionary.csv"))
# Pull all UniProt accessions annotated as keratin (Krt* gene symbol).
# The Protein column can be a semicolon-separated group, so split before matching.
keratin_ids <- protein_dict %>%
filter(str_detect(Gene, "^Krt") & !str_detect(Gene, "^Krtcap")) %>% # drop Krtcap2 (not a keratin contaminant)
pull(Protein) %>%
str_split(";") %>%
unlist() %>%
unique()
message("Keratin accessions to filter: ", length(keratin_ids))
# Match against ProteinName by splitting protein groups the same way,
# so a group is dropped if ANY member is a keratin.
msstats_input_filt <- msstats_input_filt %>%
filter(!map_lgl(str_split(ProteinName, ";"),
~ any(.x %in% keratin_ids)))
message("Rows after keratin filter: ", nrow(msstats_input_filt))
message("Proteins after keratin filter: ", n_distinct(msstats_input_filt$ProteinName))Summarises fragment intensities to protein level with TMP, median normalisation.
INFO [2026-04-29 22:26:59] ** Features with one or two measurements across runs are removed.
INFO [2026-04-29 22:27:00] ** Fractionation handled.
INFO [2026-04-29 22:27:47] ** Updated quantification data to make balanced design. Missing values are marked by NA
INFO [2026-04-29 22:28:47] ** Log2 intensities under cutoff = -2.1513 were considered as censored missing values.
INFO [2026-04-29 22:28:47] ** Log2 intensities = NA were considered as censored missing values.
INFO [2026-04-30 00:14:06] ** Flag uninformative feature and outliers by feature selection algorithm.
INFO [2026-04-30 00:14:09]
# proteins: 8890
# peptides per protein: 1-629
# features per peptide: 1-6
INFO [2026-04-30 00:14:09] Some proteins have only one feature:
A0A0G2JET4,
A6X935,
D3Z7A4;K7N6U2;K7N6Y8;K7N754;L7N2A7,
P35991,
Q64445 ...
INFO [2026-04-30 00:14:09]
hApp hAppNLGF hAppNLGF_FIRE hApp_FIRE
# runs 4 4 4 4
# bioreplicates 4 4 4 4
# tech. replicates 1 1 1 1
INFO [2026-04-30 00:14:12] Some features are completely missing in at least one condition:
AAAAAAAAQM[Oxidation (M)]HTK_2_b4_1,
AAAAAAAAQM[Oxidation (M)]HTK_2_b5_1,
AAAAAAAAQM[Oxidation (M)]HTK_2_y8_1,
AAAADGEPLHNEEER_2_y10_1,
AAAAVVEFQR_2_y4_1 ...
INFO [2026-04-30 00:14:12] == Start the summarization per subplot...
INFO [2026-04-30 00:14:12] ** Filtered out uninformative features and outliers.
|
| | 0%
|
| | 1%
|
|= | 1%
|
|= | 2%
|
|== | 2%
|
|== | 3%
|
|== | 4%
|
|=== | 4%
|
|=== | 5%
|
|==== | 5%
|
|==== | 6%
|
|===== | 6%
|
|===== | 7%
|
|===== | 8%
|
|====== | 8%
|
|====== | 9%
|
|======= | 9%
|
|======= | 10%
|
|======= | 11%
|
|======== | 11%
|
|======== | 12%
|
|========= | 12%
|
|========= | 13%
|
|========= | 14%
|
|========== | 14%
|
|========== | 15%
|
|=========== | 15%
|
|=========== | 16%
|
|============ | 16%
|
|============ | 17%
|
|============ | 18%
|
|============= | 18%
|
|============= | 19%
|
|============== | 19%
|
|============== | 20%
|
|============== | 21%
|
|=============== | 21%
|
|=============== | 22%
|
|================ | 22%
|
|================ | 23%
|
|================ | 24%
|
|================= | 24%
|
|================= | 25%
|
|================== | 25%
|
|================== | 26%
|
|=================== | 26%
|
|=================== | 27%
|
|=================== | 28%
|
|==================== | 28%
|
|==================== | 29%
|
|===================== | 29%
|
|===================== | 30%
|
|===================== | 31%
|
|====================== | 31%
|
|====================== | 32%
|
|======================= | 32%
|
|======================= | 33%
|
|======================= | 34%
|
|======================== | 34%
|
|======================== | 35%
|
|========================= | 35%
|
|========================= | 36%
|
|========================== | 36%
|
|========================== | 37%
|
|========================== | 38%
|
|=========================== | 38%
|
|=========================== | 39%
|
|============================ | 39%
|
|============================ | 40%
|
|============================ | 41%
|
|============================= | 41%
|
|============================= | 42%
|
|============================== | 42%
|
|============================== | 43%
|
|============================== | 44%
|
|=============================== | 44%
|
|=============================== | 45%
|
|================================ | 45%
|
|================================ | 46%
|
|================================= | 46%
|
|================================= | 47%
|
|================================= | 48%
|
|================================== | 48%
|
|================================== | 49%
|
|=================================== | 49%
|
|=================================== | 50%
|
|=================================== | 51%
|
|==================================== | 51%
|
|==================================== | 52%
|
|===================================== | 52%
|
|===================================== | 53%
|
|===================================== | 54%
|
|====================================== | 54%
|
|====================================== | 55%
|
|======================================= | 55%
|
|======================================= | 56%
|
|======================================== | 56%
|
|======================================== | 57%
|
|======================================== | 58%
|
|========================================= | 58%
|
|========================================= | 59%
|
|========================================== | 59%
|
|========================================== | 60%
|
|========================================== | 61%
|
|=========================================== | 61%
|
|=========================================== | 62%
|
|============================================ | 62%
|
|============================================ | 63%
|
|============================================ | 64%
|
|============================================= | 64%
|
|============================================= | 65%
|
|============================================== | 65%
|
|============================================== | 66%
|
|=============================================== | 66%
|
|=============================================== | 67%
|
|=============================================== | 68%
|
|================================================ | 68%
|
|================================================ | 69%
|
|================================================= | 69%
|
|================================================= | 70%
|
|================================================= | 71%
|
|================================================== | 71%
|
|================================================== | 72%
|
|=================================================== | 72%
|
|=================================================== | 73%
|
|=================================================== | 74%
|
|==================================================== | 74%
|
|==================================================== | 75%
|
|===================================================== | 75%
|
|===================================================== | 76%
|
|====================================================== | 76%
|
|====================================================== | 77%
|
|====================================================== | 78%
|
|======================================================= | 78%
|
|======================================================= | 79%
|
|======================================================== | 79%
|
|======================================================== | 80%
|
|======================================================== | 81%
|
|========================================================= | 81%
|
|========================================================= | 82%
|
|========================================================== | 82%
|
|========================================================== | 83%
|
|========================================================== | 84%
|
|=========================================================== | 84%
|
|=========================================================== | 85%
|
|============================================================ | 85%
|
|============================================================ | 86%
|
|============================================================= | 86%
|
|============================================================= | 87%
|
|============================================================= | 88%
|
|============================================================== | 88%
|
|============================================================== | 89%
|
|=============================================================== | 89%
|
|=============================================================== | 90%
|
|=============================================================== | 91%
|
|================================================================ | 91%
|
|================================================================ | 92%
|
|================================================================= | 92%
|
|================================================================= | 93%
|
|================================================================= | 94%
|
|================================================================== | 94%
|
|================================================================== | 95%
|
|=================================================================== | 95%
|
|=================================================================== | 96%
|
|==================================================================== | 96%
|
|==================================================================== | 97%
|
|==================================================================== | 98%
|
|===================================================================== | 98%
|
|===================================================================== | 99%
|
|======================================================================| 99%
|
|======================================================================| 100%
INFO [2026-04-30 00:30:47] == Summarization is done.MSstats sorts conditions alphabetically — confirm the order before building the contrast matrix.
Condition levels (alphabetical order used in contrast matrix):[1] "hApp" "hApp_FIRE" "hAppNLGF" "hAppNLGF_FIRE"Pairwise comparisons (numerator vs denominator):
| Contrast | Question |
|---|---|
| hAppNLGF vs hApp | NLGF mutation effect in microglia-containing condition |
| hApp_FIRE vs hApp | Microglia depletion effect on hApp background |
| hAppNLGF_FIRE vs hAppNLGF | Microglia depletion effect on NLGF background |
| hAppNLGF_FIRE vs hApp_FIRE | NLGF mutation effect in microglia-depleted condition |
The 2×2 design (APP genotype × microglia presence) also supports an interaction contrast, which directly tests whether the NLGF effect depends on microglia presence — i.e. proteins where microglia change how the proteome reacts to amyloid pathology. This is a single statistical test (one p-value per protein) and is the proper way to identify “microglia-amplified APP-NLGF responses” rather than informally comparing two pairwise contrasts.
Interaction = (hAppNLGF − hApp) − (hAppNLGF_FIRE − hApp_FIRE)
# Helper: builds a named contrast vector given the full condition_levels vector
make_contrast <- function(numerator, denominator, levels) {
v <- setNames(rep(0L, length(levels)), levels)
v[numerator] <- 1L
v[denominator] <- -1L
v
}
# Helper: arbitrary linear combination across conditions
make_linear_contrast <- function(weights, levels) {
v <- setNames(rep(0, length(levels)), levels)
v[names(weights)] <- weights
stopifnot(abs(sum(v)) < 1e-9) # contrast must sum to zero
v
}
# APP × Microglia interaction:
# +1 hApp (WT × micro+)
# -1 hAppNLGF (NLGF × micro+)
# -1 hApp_FIRE (WT × micro-)
# +1 hAppNLGF_FIRE (NLGF × micro-)
# Equivalently: (hAppNLGF - hApp) - (hAppNLGF_FIRE - hApp_FIRE), negated so a
# positive log2FC means NLGF effect is *stronger* with microglia present.
interaction_weights <- c(
hApp = -1,
hAppNLGF = 1,
hApp_FIRE = 1,
hAppNLGF_FIRE = -1
)
comparison_matrix <- rbind(
make_contrast("hAppNLGF", "hApp", condition_levels),
make_contrast("hApp_FIRE", "hApp", condition_levels),
make_contrast("hAppNLGF_FIRE", "hAppNLGF", condition_levels),
make_contrast("hAppNLGF_FIRE", "hApp_FIRE", condition_levels),
make_linear_contrast(interaction_weights, condition_levels)
)
rownames(comparison_matrix) <- c(
"hAppNLGF_vs_hApp",
"hApp_FIRE_vs_hApp",
"hAppNLGF_FIRE_vs_hAppNLGF",
"hAppNLGF_FIRE_vs_hApp_FIRE",
"Interaction_APP_x_Microglia"
)
print(comparison_matrix) hApp hApp_FIRE hAppNLGF hAppNLGF_FIRE
hAppNLGF_vs_hApp -1 0 1 0
hApp_FIRE_vs_hApp -1 1 0 0
hAppNLGF_FIRE_vs_hAppNLGF 0 0 -1 1
hAppNLGF_FIRE_vs_hApp_FIRE 0 -1 0 1
Interaction_APP_x_Microglia -1 1 1 -1Linear mixed-effects model across all proteins.
INFO [2026-04-30 00:31:12] == Start to test and get inference in whole plot ...
|
| | 0%
|
| | 1%
|
|= | 1%
|
|= | 2%
|
|== | 2%
|
|== | 3%
|
|== | 4%
|
|=== | 4%
|
|=== | 5%
|
|==== | 5%
|
|==== | 6%
|
|===== | 6%
|
|===== | 7%
|
|===== | 8%
|
|====== | 8%
|
|====== | 9%
|
|======= | 9%
|
|======= | 10%
|
|======= | 11%
|
|======== | 11%
|
|======== | 12%
|
|========= | 12%
|
|========= | 13%
|
|========= | 14%
|
|========== | 14%
|
|========== | 15%
|
|=========== | 15%
|
|=========== | 16%
|
|============ | 16%
|
|============ | 17%
|
|============ | 18%
|
|============= | 18%
|
|============= | 19%
|
|============== | 19%
|
|============== | 20%
|
|============== | 21%
|
|=============== | 21%
|
|=============== | 22%
|
|================ | 22%
|
|================ | 23%
|
|================ | 24%
|
|================= | 24%
|
|================= | 25%
|
|================== | 25%
|
|================== | 26%
|
|=================== | 26%
|
|=================== | 27%
|
|=================== | 28%
|
|==================== | 28%
|
|==================== | 29%
|
|===================== | 29%
|
|===================== | 30%
|
|===================== | 31%
|
|====================== | 31%
|
|====================== | 32%
|
|======================= | 32%
|
|======================= | 33%
|
|======================= | 34%
|
|======================== | 34%
|
|======================== | 35%
|
|========================= | 35%
|
|========================= | 36%
|
|========================== | 36%
|
|========================== | 37%
|
|========================== | 38%
|
|=========================== | 38%
|
|=========================== | 39%
|
|============================ | 39%
|
|============================ | 40%
|
|============================ | 41%
|
|============================= | 41%
|
|============================= | 42%
|
|============================== | 42%
|
|============================== | 43%
|
|============================== | 44%
|
|=============================== | 44%
|
|=============================== | 45%
|
|================================ | 45%
|
|================================ | 46%
|
|================================= | 46%
|
|================================= | 47%
|
|================================= | 48%
|
|================================== | 48%
|
|================================== | 49%
|
|=================================== | 49%
|
|=================================== | 50%
|
|=================================== | 51%
|
|==================================== | 51%
|
|==================================== | 52%
|
|===================================== | 52%
|
|===================================== | 53%
|
|===================================== | 54%
|
|====================================== | 54%
|
|====================================== | 55%
|
|======================================= | 55%
|
|======================================= | 56%
|
|======================================== | 56%
|
|======================================== | 57%
|
|======================================== | 58%
|
|========================================= | 58%
|
|========================================= | 59%
|
|========================================== | 59%
|
|========================================== | 60%
|
|========================================== | 61%
|
|=========================================== | 61%
|
|=========================================== | 62%
|
|============================================ | 62%
|
|============================================ | 63%
|
|============================================ | 64%
|
|============================================= | 64%
|
|============================================= | 65%
|
|============================================== | 65%
|
|============================================== | 66%
|
|=============================================== | 66%
|
|=============================================== | 67%
|
|=============================================== | 68%
|
|================================================ | 68%
|
|================================================ | 69%
|
|================================================= | 69%
|
|================================================= | 70%
|
|================================================= | 71%
|
|================================================== | 71%
|
|================================================== | 72%
|
|=================================================== | 72%
|
|=================================================== | 73%
|
|=================================================== | 74%
|
|==================================================== | 74%
|
|==================================================== | 75%
|
|===================================================== | 75%
|
|===================================================== | 76%
|
|====================================================== | 76%
|
|====================================================== | 77%
|
|====================================================== | 78%
|
|======================================================= | 78%
|
|======================================================= | 79%
|
|======================================================== | 79%
|
|======================================================== | 80%
|
|======================================================== | 81%
|
|========================================================= | 81%
|
|========================================================= | 82%
|
|========================================================== | 82%
|
|========================================================== | 83%
|
|========================================================== | 84%
|
|=========================================================== | 84%
|
|=========================================================== | 85%
|
|============================================================ | 85%
|
|============================================================ | 86%
|
|============================================================= | 86%
|
|============================================================= | 87%
|
|============================================================= | 88%
|
|============================================================== | 88%
|
|============================================================== | 89%
|
|=============================================================== | 89%
|
|=============================================================== | 90%
|
|=============================================================== | 91%
|
|================================================================ | 91%
|
|================================================================ | 92%
|
|================================================================= | 92%
|
|================================================================= | 93%
|
|================================================================= | 94%
|
|================================================================== | 94%
|
|================================================================== | 95%
|
|=================================================================== | 95%
|
|=================================================================== | 96%
|
|==================================================================== | 96%
|
|==================================================================== | 97%
|
|==================================================================== | 98%
|
|===================================================================== | 98%
|
|===================================================================== | 99%
|
|======================================================================| 99%
|
|======================================================================| 100%
INFO [2026-04-30 00:33:04] == Comparisons for all proteins are done.Total protein-comparison rows: 44450 Significant (adj.pvalue < 0.05): 326 Results saved to: /nemo/lab/destrooperb/home/shared/zanettc/giulia_proteomics/bulk_proteomics/results/run4/Full_GroupComparison_Results.csv ---
title: "MSstats — Differential Abundance Analysis"
---
## Libraries
```{r setup, include=TRUE, message=FALSE}
library(data.table)
library(MSstats)
library(qs2)
library(dplyr)
library(ggplot2)
library(ggrepel)
library(tidyverse)
library(readr)
```
## Set directories
```{r}
base_dir <- "/nemo/lab/destrooperb/home/shared/zanettc/giulia_proteomics/bulk_proteomics"
run_num <- "run4"
results_dir <- file.path(base_dir, "results", run_num)
objects_dir <- file.path(base_dir, "data", "processed", run_num)
input_dir <- file.path(base_dir, "input")
dir.create(results_dir, recursive = TRUE, showWarnings = FALSE)
dir.create(objects_dir, recursive = TRUE, showWarnings = FALSE)
```
## Build annotation from study design
All 16 samples passed QC. R.Condition = "Not Defined" in the raw file because the study
design was not configured inside Spectronaut — we define it here from the known design.
```{r}
msstats_annotation <- data.table(
Run = paste0("GA_", 1:16),
Condition = rep(c("hApp", "hAppNLGF", "hAppNLGF_FIRE", "hApp_FIRE"), each = 4),
BioReplicate = 1:16
)
msstats_annotation
fwrite(msstats_annotation, file.path(objects_dir, "msstats_annotation.csv"))
```
## Load raw MSstats Spectronaut proteome report
Only the columns required by MSstats are read — this keeps memory manageable on the 8.4 GB file.
```{r}
msstats_cols <- c(
"R.Condition", "R.FileName", "R.Replicate",
"PG.ProteinGroups", "PG.ProteinAccessions",
"PEP.StrippedSequence",
"EG.ModifiedSequence", "EG.PrecursorId", "EG.Qvalue",
"FG.Charge", "FG.Id",
"F.FrgIon", "F.Charge", "F.FrgLossType", "F.FrgNum", "F.FrgType",
"F.NormalizedPeakArea", "F.PeakArea", "F.ExcludedFromQuantification"
)
cat("Reading Spectronaut MSstats report...\n")
raw_spectronaut <- fread(
file.path(input_dir, "20260423_154110_PCF000612_GA_MSStats_Report-Proteome.csv"),
select = msstats_cols
)
cat("Rows loaded:", nrow(raw_spectronaut), "\n")
qs_save(raw_spectronaut, file.path(objects_dir, "msstats_report.qs2"))
```
## Reload from .qs2 (skip the read step on re-runs)
```{r}
#raw_spectronaut <- qs_read(file.path(objects_dir, "msstats_report.qs2"))
```
## Overwrite R.Condition and R.Replicate with study design values
SpectronauttoMSstatsFormat uses R.Condition internally before merging with the annotation.
We replace "Not Defined" with the correct values from the annotation to avoid any issues.
```{r}
raw_spectronaut[msstats_annotation,
on = c("R.FileName" = "Run"),
`:=`(R.Condition = i.Condition,
R.Replicate = i.BioReplicate)]
cat("Condition distribution after fix:\n")
print(raw_spectronaut[, .N, by = .(R.Condition, R.FileName)][order(R.Condition, R.FileName)])
```
## Amyloid-β tryptic peptide verification
Confirms that hApp and hAppNLGF conditions have detectable Aβ signal.
In microglia, signal reflects phagocytosed / secreted Aβ rather than endogenous APP expression.
```{r}
arctic_peptide <- "LVFFAGDVGSNK"
normal_peptide <- "LVFFAEDVGSNK"
peptide_both <- "DAEFRHDSGYEVHHQK"
target_peptide <- "LVFFAEDVGSNK" # Conserved Aβ tryptic peptide (mouse and human APP)
ab_check <- raw_spectronaut %>%
filter(PEP.StrippedSequence == target_peptide) %>%
group_by(R.FileName, R.Condition) %>%
summarise(Total_Intensity = sum(F.PeakArea, na.rm = TRUE), .groups = "drop") %>%
mutate(Log2_Intensity = log2(Total_Intensity + 1)) %>%
# Extract numeric sample ID for proper sorting
mutate(Sample_Num = as.numeric(gsub("GA_", "", R.FileName))) %>%
arrange(R.Condition, Sample_Num) %>%
mutate(R.FileName = factor(R.FileName, levels = unique(R.FileName)))
if (nrow(ab_check) == 0) {
cat("Aβ peptide (", target_peptide, ") not detected in any sample.\n")
} else {
cat("Aβ peptide detected in", nrow(ab_check), "sample(s):\n")
print(ab_check)
}
p_ab <- ggplot(ab_check, aes(x = R.FileName, y = Log2_Intensity, fill = R.Condition)) +
geom_bar(stat = "identity") +
scale_fill_manual(values = c(
"hApp" = "#1b9e77", "hAppNLGF" = "#d95f02",
"hAppNLGF_FIRE" = "#7570b3", "hApp_FIRE" = "#e7298a"
)) +
theme_minimal() +
theme(axis.text.x = element_text(angle = 45, hjust = 1)) +
labs(title = "Aβ Tryptic Peptide Signal per Sample",
subtitle = paste0("Peptide: ", target_peptide),
x = "Sample", y = "Log2 Intensity")
ggsave(file.path(results_dir, "AB_Peptide_Verification.pdf"), p_ab, width = 10, height = 6)
print(p_ab)
```
## Convert to MSstats input format
Feature-level filtering: Qvalue < 0.01, remove features with too few measurements.
```{r}
msstats_input <- suppressWarnings(SpectronauttoMSstatsFormat(
raw_spectronaut,
annotation = msstats_annotation,
filter_with_Qvalue = TRUE,
qvalue_cutoff = 0.01,
fewMeasurements = "remove",
removeProtein_with1Feature = FALSE
))
cat("MSstats input rows:", nrow(msstats_input), "\n")
cat("Unique proteins: ", uniqueN(msstats_input$ProteinName), "\n")
qs_save(msstats_input, file.path(objects_dir, "msstats_input.qs2"))
```
## Feature completeness filter
Keeps features observed in ≥50% of runs within at least one condition.
This removes near-absent features that destabilise the linear model.
This is the same as saying proteins must be present in at least 2 samples out
of 3 for at least one condition.
```{r reload, eval=FALSE}
# Manual resume only — not evaluated during render. msstats_input is already
# in memory from the chunk above; this line lets you skip rebuilding it on a re-run.
msstats_input <- qs_read(file.path(objects_dir, "msstats_input.qs2"))
```
```{r}
msstats_input_filt <- as.data.frame(msstats_input) %>%
group_by(ProteinName, PeptideSequence, PrecursorCharge, FragmentIon, ProductCharge, Condition) %>%
summarise(
n_obs = sum(!is.na(Intensity) & Intensity > 0),
n_total = n(),
prop = n_obs / n_total,
.groups = "drop"
) %>%
group_by(ProteinName, PeptideSequence, PrecursorCharge, FragmentIon, ProductCharge) %>%
filter(any(prop >= 0.5)) %>%
ungroup() %>%
inner_join(
as.data.frame(msstats_input),
by = c("ProteinName", "PeptideSequence", "PrecursorCharge",
"FragmentIon", "ProductCharge", "Condition")
)
message("Rows before completeness filter: ", nrow(msstats_input))
message("Rows after completeness filter: ", nrow(msstats_input_filt))
message("Proteins retained: ", uniqueN(msstats_input_filt$ProteinName))
table(
Value_Type = case_when(
is.na(msstats_input$Intensity) ~ "True NAs",
msstats_input$Intensity == 0 ~ "Exact Zeros (Filtered by Q-value)",
msstats_input$Intensity > 0 ~ "Valid Measurements"
)
)
```
## (Optional) Keratin protein filter
The MSstats report contains only UniProt accessions in ProteinName — no gene symbols.
To filter keratins, build a lookup from UniProt or biomaRt and enable the block below.
```{r}
## Keratin protein filter
protein_dict <- read_csv(file.path(base_dir, "data", "processed", "run1", "protein_dictionary.csv"))
# Pull all UniProt accessions annotated as keratin (Krt* gene symbol).
# The Protein column can be a semicolon-separated group, so split before matching.
keratin_ids <- protein_dict %>%
filter(str_detect(Gene, "^Krt") & !str_detect(Gene, "^Krtcap")) %>% # drop Krtcap2 (not a keratin contaminant)
pull(Protein) %>%
str_split(";") %>%
unlist() %>%
unique()
message("Keratin accessions to filter: ", length(keratin_ids))
# Match against ProteinName by splitting protein groups the same way,
# so a group is dropped if ANY member is a keratin.
msstats_input_filt <- msstats_input_filt %>%
filter(!map_lgl(str_split(ProteinName, ";"),
~ any(.x %in% keratin_ids)))
message("Rows after keratin filter: ", nrow(msstats_input_filt))
message("Proteins after keratin filter: ", n_distinct(msstats_input_filt$ProteinName))
```
## Data summarisation and normalisation
Summarises fragment intensities to protein level with TMP, median normalisation.
```{r}
processed_data <- suppressWarnings(dataProcess(
msstats_input_filt,
normalization = "equalizeMedians",
summaryMethod = "TMP",
censoredInt = 'NA',
MBimpute = TRUE,
featureSubset = "highQuality",
remove_uninformative_feature_outlier = TRUE
))
qs_save(processed_data, file.path(objects_dir, "processed_msstats_data.qs2"))
```
## Reload processed data (if re-running from this point)
```{r}
processed_data <- qs_read(file.path(objects_dir, "processed_msstats_data.qs2"))
```
## Verify condition levels
MSstats sorts conditions alphabetically — confirm the order before building the contrast matrix.
```{r}
condition_levels <- levels(processed_data$ProteinLevelData$GROUP)
cat("Condition levels (alphabetical order used in contrast matrix):\n")
print(condition_levels)
# Expected: "hApp" "hApp_FIRE" "hAppNLGF" "hAppNLG_FIRE"
# (underscore sorts after capital letters in most R locales,
# but verify with the output above before proceeding)
```
## Build contrast matrix
Pairwise comparisons (numerator vs denominator):
| Contrast | Question |
|---|---|
| hAppNLGF vs hApp | NLGF mutation effect in microglia-containing condition |
| hApp_FIRE vs hApp | Microglia depletion effect on hApp background |
| hAppNLGF_FIRE vs hAppNLGF | Microglia depletion effect on NLGF background |
| hAppNLGF_FIRE vs hApp_FIRE | NLGF mutation effect in microglia-depleted condition |
The 2×2 design (APP genotype × microglia presence) also supports an **interaction contrast**, which directly tests whether the NLGF effect depends on microglia presence — i.e. proteins where microglia *change how the proteome reacts* to amyloid pathology. This is a single statistical test (one p-value per protein) and is the proper way to identify "microglia-amplified APP-NLGF responses" rather than informally comparing two pairwise contrasts.
Interaction = `(hAppNLGF − hApp) − (hAppNLGF_FIRE − hApp_FIRE)`
```{r}
# Helper: builds a named contrast vector given the full condition_levels vector
make_contrast <- function(numerator, denominator, levels) {
v <- setNames(rep(0L, length(levels)), levels)
v[numerator] <- 1L
v[denominator] <- -1L
v
}
# Helper: arbitrary linear combination across conditions
make_linear_contrast <- function(weights, levels) {
v <- setNames(rep(0, length(levels)), levels)
v[names(weights)] <- weights
stopifnot(abs(sum(v)) < 1e-9) # contrast must sum to zero
v
}
# APP × Microglia interaction:
# +1 hApp (WT × micro+)
# -1 hAppNLGF (NLGF × micro+)
# -1 hApp_FIRE (WT × micro-)
# +1 hAppNLGF_FIRE (NLGF × micro-)
# Equivalently: (hAppNLGF - hApp) - (hAppNLGF_FIRE - hApp_FIRE), negated so a
# positive log2FC means NLGF effect is *stronger* with microglia present.
interaction_weights <- c(
hApp = -1,
hAppNLGF = 1,
hApp_FIRE = 1,
hAppNLGF_FIRE = -1
)
comparison_matrix <- rbind(
make_contrast("hAppNLGF", "hApp", condition_levels),
make_contrast("hApp_FIRE", "hApp", condition_levels),
make_contrast("hAppNLGF_FIRE", "hAppNLGF", condition_levels),
make_contrast("hAppNLGF_FIRE", "hApp_FIRE", condition_levels),
make_linear_contrast(interaction_weights, condition_levels)
)
rownames(comparison_matrix) <- c(
"hAppNLGF_vs_hApp",
"hApp_FIRE_vs_hApp",
"hAppNLGF_FIRE_vs_hAppNLGF",
"hAppNLGF_FIRE_vs_hApp_FIRE",
"Interaction_APP_x_Microglia"
)
print(comparison_matrix)
```
## Statistical modelling
Linear mixed-effects model across all proteins.
```{r}
test_results <- suppressWarnings(groupComparison(contrast.matrix = comparison_matrix, data = processed_data))
qs_save(test_results, file.path(objects_dir, "comparison_results.qs2"))
res_df <- as.data.table(test_results$ComparisonResult)
cat("Total protein-comparison rows:", nrow(res_df), "\n")
cat("Significant (adj.pvalue < 0.05):",
nrow(res_df[adj.pvalue < 0.05, ]), "\n")
fwrite(res_df, file.path(results_dir, "Full_GroupComparison_Results.csv"))
cat("Results saved to:", file.path(results_dir, "Full_GroupComparison_Results.csv"), "\n")
```
## Quick summary of results
```{r}
res_df %>%
filter(!is.na(adj.pvalue)) %>%
group_by(Label) %>%
summarise(
n_tested = n(),
n_sig = sum(adj.pvalue < 0.05, na.rm = TRUE),
n_up = sum(adj.pvalue < 0.05 & log2FC > 0, na.rm = TRUE),
n_down = sum(adj.pvalue < 0.05 & log2FC < 0, na.rm = TRUE),
.groups = "drop"
) %>%
arrange(Label)
```
## Reload comparison results (if re-running from this point)
```{r}
#test_results <- qs_read(file.path(objects_dir, "comparison_results.qs2"))
#res_df <- as.data.table(test_results$ComparisonResult)
```